Posts Search
23 Results
How to make your research software citable
Making your research software citable means you get credit for that work and others can build on it with confidence. This practical guide will show you how.
N8CIR Digital Research Internships 2026: Applications Now Open
Applications are now open for the 2026 N8 Digital Research Internships, offering undergraduate students the chance to work on real research projects with our team this summer.
Research Storage Pricing Update – August 2026
From 1 August 2026, pricing for the University’s Research Data Storage service will change.
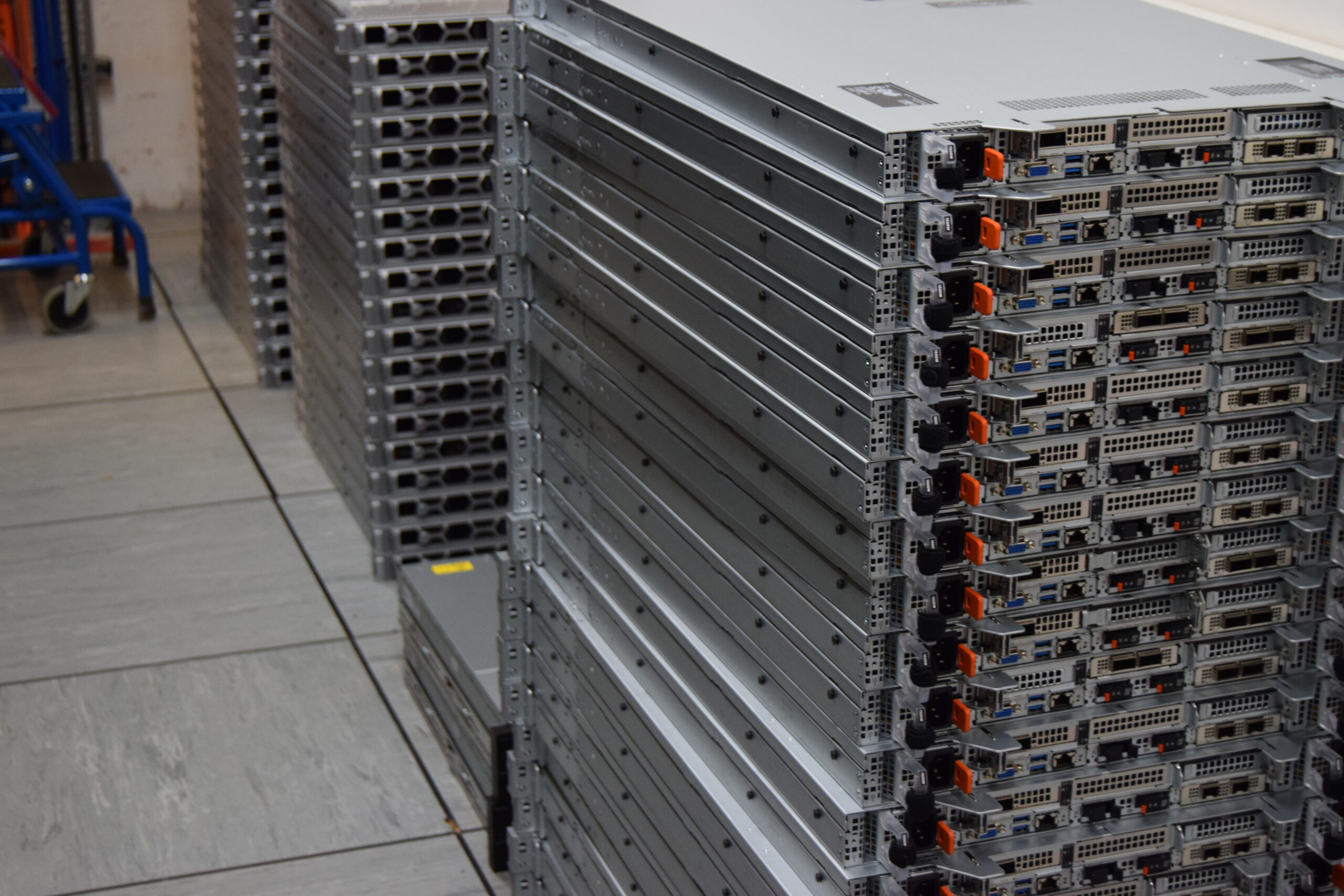
Calder Progress Update – March 2026
Calder, the University’s next HPC cluster, continues to take shape as a steady stream of hardware arrives.
Call open for RSE summer internship project proposals
Got a research idea gathering dust? Find out how an RSE summer internship could help breathe life into it.
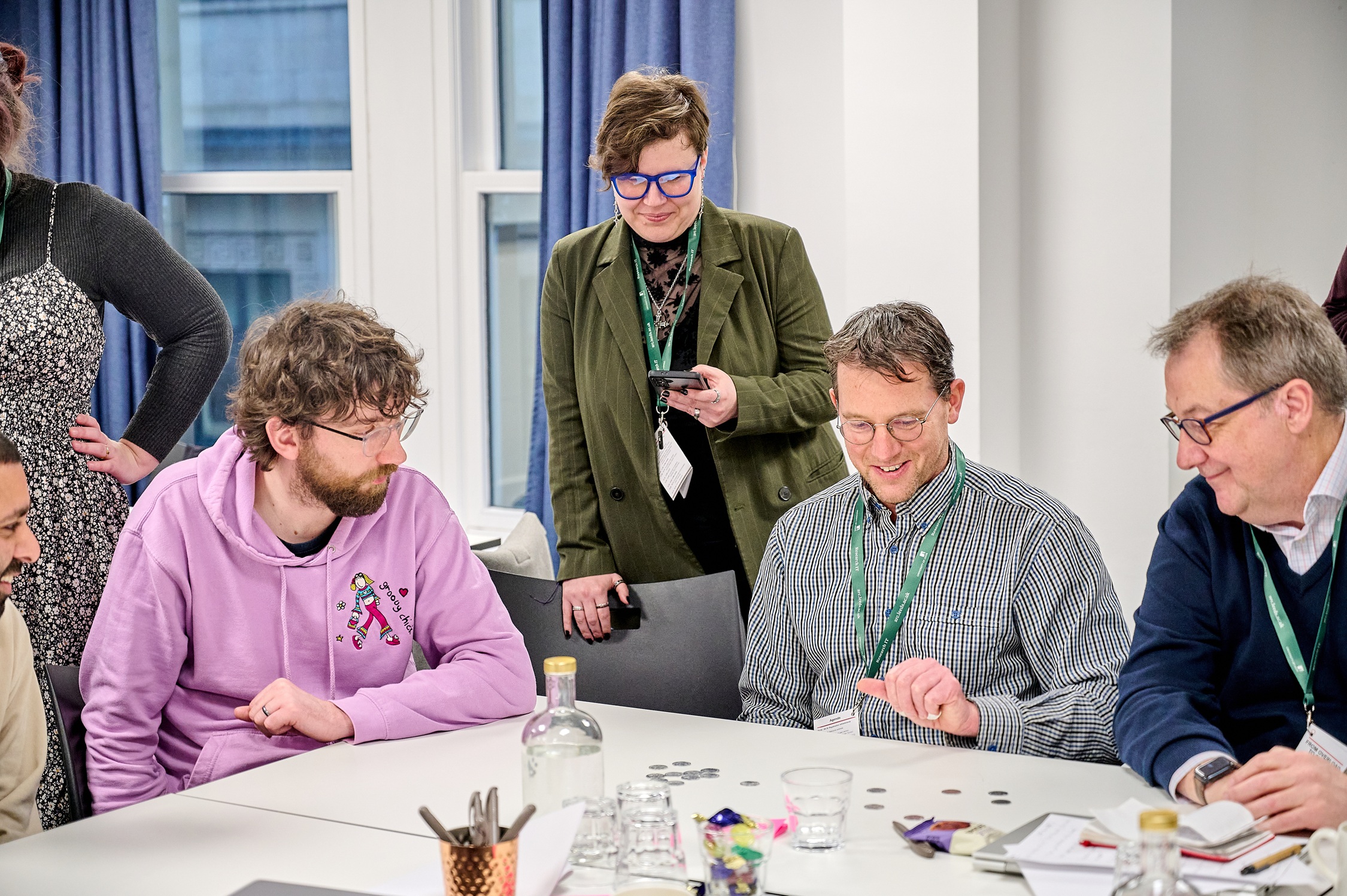
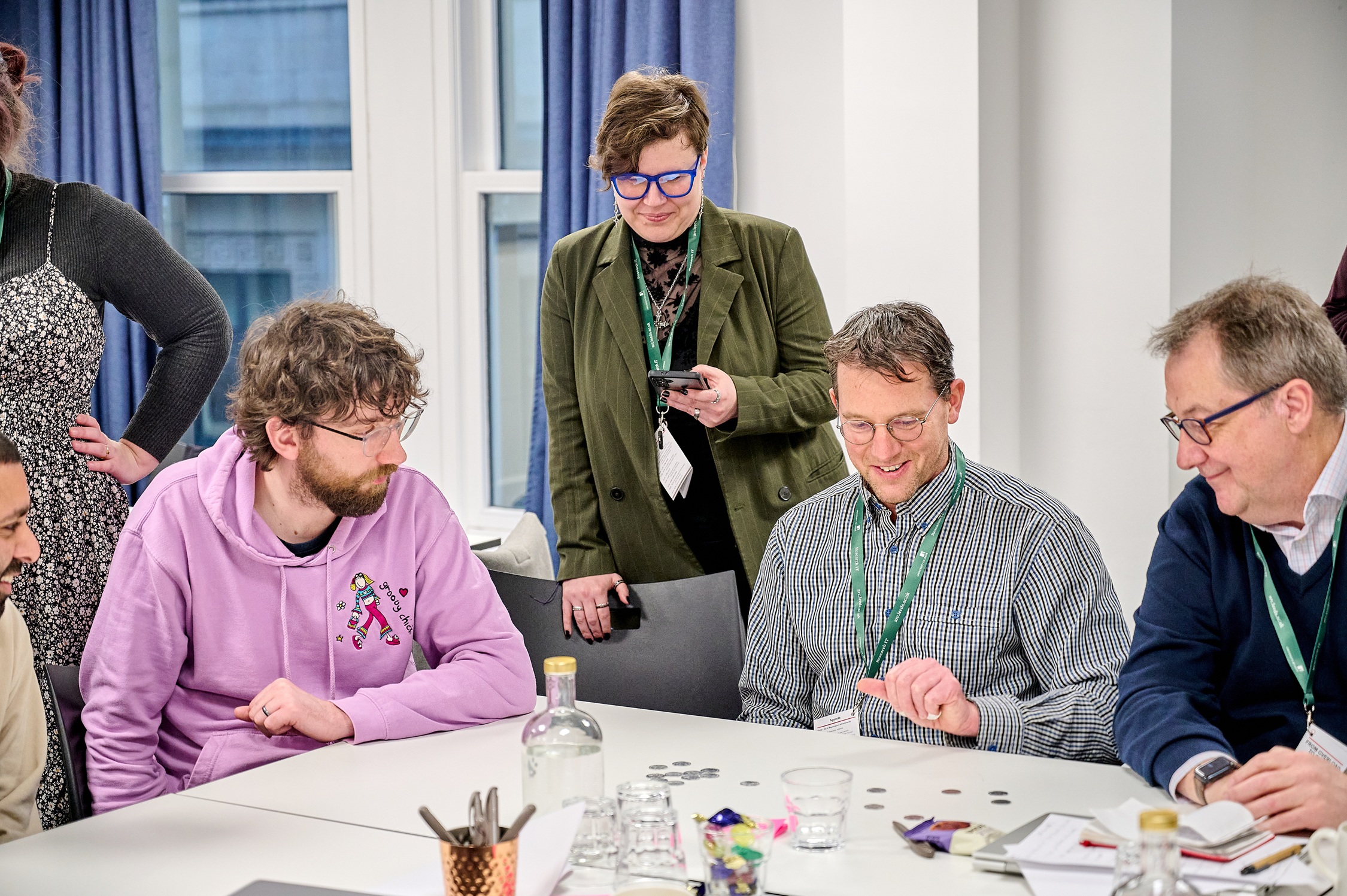
From Overload to Clarity
Key learnings from our recent RSE learning event, From Overload to Clarity - a day spent exploring user-centred service design and systems thinking.
Using GenAI to Build Research Software: A Conversation About What Works
Does GenAI have a place in research software development? We chat to Leeds biophysicist and self-taught developer, Daniel Rollins, about what works and what doesn't.
What if digital research infrastructure was designed with arts and humanities in mind?
Digital research infrastructure designed for science can create unexpected barriers for researchers in the arts and humanities. In this article, we explore what a more arts-focused infrastructure might look like.
Calder Unveiled: Powering Discovery with Our Next HPC System
As we move into the purchase phase of the Calder tender, we’re excited to share the first public look at the system that will power the next generation of computational research at Leeds.
1/3